Some developments- (this should be interesting to Microsoft that is basically leading player in Enterprise Solutions in R after completely acquiring Revolution R and RStudio being headed by another Microsoft alum)
- you can install R using miniconda http://continuum.io/conda-for-R
- you can run R using Jupyter notebooks
and see
http://continuum.io/blog/conda-jupyter-irkernel
“R Essentials” setup
The Anaconda team has created an “R Essentials” bundle with the IRKernel and over 80 of the most used R packages for data science, including dplyr, shiny, ggplot2, tidyr,caret and nnet.
Downloading “R Essentials” requires conda. Miniconda includes conda, Python, and a few other necessary packages, while Anaconda includes all this and over 200 of the most popularPython packages for science, math, engineering, and data analysis. Users may install all of Anaconda at once, or they may install Miniconda at first and then use conda to install any other packages they need, including any of the packages in Anaconda.
Once you have conda, you may install “R Essentials” into the current environment:
conda install -c r r-essentials
or create a new environment just for “R essentials”:
conda create -n my-r-env -c r r-essentials
Jupyter
Jupyter provides a great notebook interface to write your analysis and share it with your peers. Open a shell and run this command to start the Jupyter notebook interface in your browser:
jupyter notebook
Start a new R notebook:
You can immediately write and run R code in the notebook cells.
- Running R from within Python – yeah!
http://blog.revolutionanalytics.com/2015/09/using-r-with-jupyter-notebooks.html
Step 1: install miniConda
- Get and install miniConda for Python 3 at http://conda.pydata.org/miniconda.html
- Important: install python 3
Step 2: open an OS terminal window:
conda install -c r ipython-notebook r-irkernel
ipython notebook
4.5
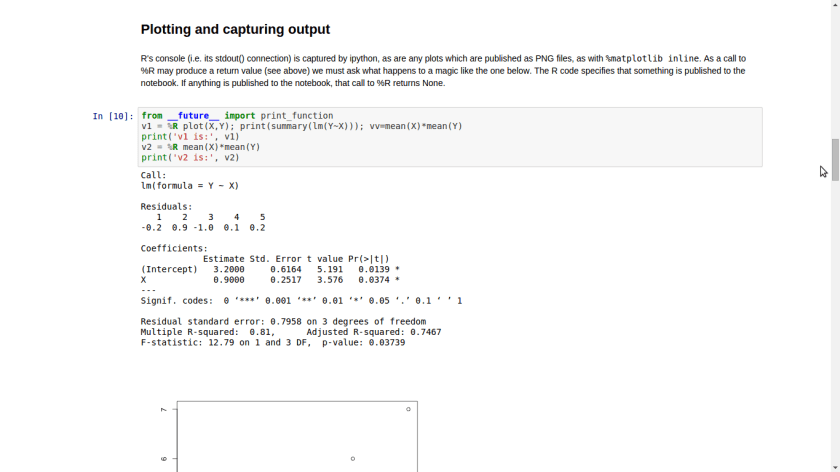
Using R Within the IPython Notebok
Using the rmagic extension, users can run R code from within the IPython Notebook. This example Notebook demonstrates this capability.
- Use docker !
https://hub.docker.com/r/jupyter/datascience-notebook/.
Jupyter Notebook Data Science Stack
What it Gives You
- Jupyter Notebook server v4.0.x
- Conda Python 3.4.x and Python 2.7.x environments
- pandas, matplotlib, scipy, seaborn, scikit-learn, scikit-image, sympy, cython, patsy, statsmodel, cloudpickle, dill, numba, bokeh pre-installed
- Conda R v3.2.x and channel
- plyr, devtools, dplyr, ggplot2, tidyr, shiny, rmarkdown, forecast, stringr, rsqlite, reshape2, nycflights13, caret, rcurl, and randomforest pre-installed
- Julia v0.3.x with Gadfly and RDatasets pre-installed
- Unprivileged user
jovyan(uid=1000, configurable, see options) in groupusers(gid=100) with ownership over/home/jovyanand/opt/conda - Options for HTTPS, password auth, and passwordless
sudo
Basic Use
The following command starts a container with the Notebook server listening for HTTP connections on port 8888 without authentication configured.
docker run -d -p 8888:8888 jupyter/datascience-notebook