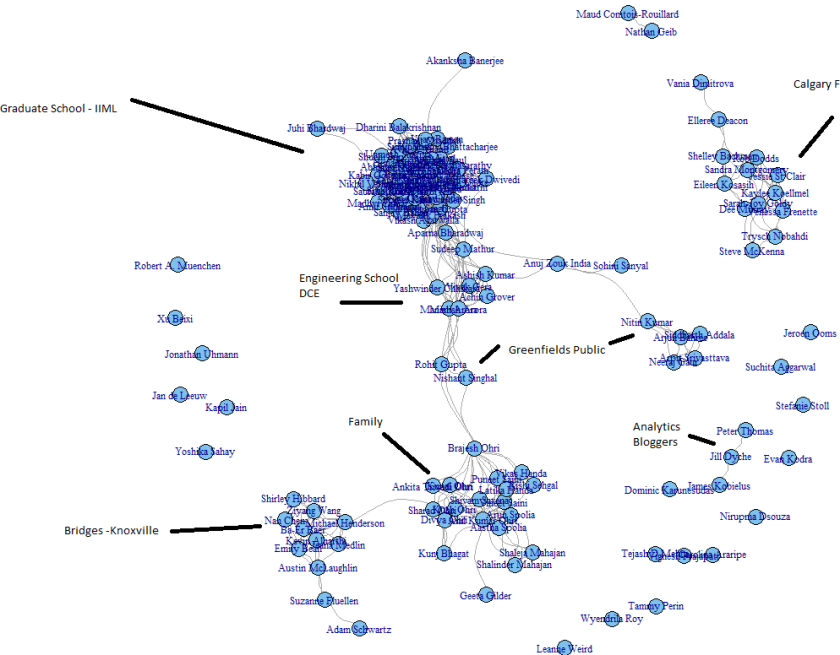
brilliant piece of why scientists write bad code
What You're Doing Is Rather Desperate
There’s a lot of discussion around why code written by self-taught “scientist programmers” rarely follows what a trained computer scientist would consider “best practice”. Here’s a recent post on the topic.
One answer: we begin with exploratory data analysis and never get around to cleaning it up.
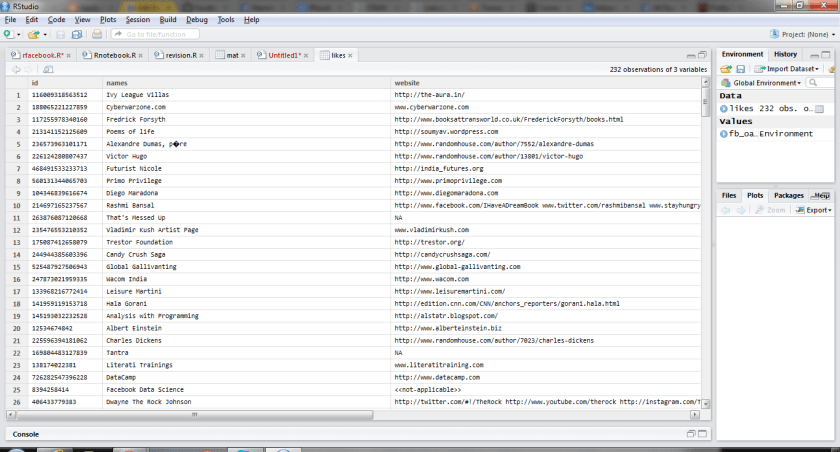
An example. For some reason, a researcher (let’s call him “Bob”) becomes interested in a particular dataset in the GEO database. So Bob opens the R console and use the GEOquery package to grab the data:
Bob is interested in the covariates and metadata associated with the experiment, which he can access using pData().
Bob discovers that pd$characteristics_ch1.2 is “age at examination”, but it’s stored as a factor. He’d like to use it as a numeric variable. So he sets about figuring out how to do the conversion.
Three levels of nested methods. Ugly. However, it works, so Bob moves to…
View original post 34 more words